Time series forecasting is a necessary scientific and business problem and as such has also seen numerous innovation recently with using deep learning based models along with the classical methods. A very important difference between classical methods like ARIMA and novel deep learning methods is the next.
Probabilistic Forecasting
Typically, classical methods are fitted on every time series in a dataset individually. These are also known as “single” or “local” methods. Nevertheless, when coping with a considerable amount of time series for some applications, it is helpful to coach a “global” model on all available time series, which enables the model to learn latent representations from many various sources.
Some classical methods are point-valued (meaning, they only output a single value per time step) and models are trained by minimizing an L2 or L1 style of loss with respect to the bottom truth data. Nevertheless, since forecasts are sometimes utilized in some real-world decision making pipeline, even with humans within the loop, it’s rather more helpful to supply the uncertainties of predictions. This can be called “probabilistic forecasting”, versus “point forecasting”. This entails modeling a probabilistic distribution, from which one can sample.
So in brief, somewhat than training local point forecasting models, we hope to coach global probabilistic models. Deep learning is an awesome fit for this, as neural networks can learn representations from several related time series in addition to model the uncertainty of the info.
It is not uncommon within the probabilistic setting to learn the longer term parameters of some chosen parametric distribution, like Gaussian or Student-T; or learn the conditional quantile function; or use the framework of Conformal Prediction adapted to the time series setting. The alternative of method doesn’t affect the modeling aspect and thus could be typically considered yet one more hyperparameter. One can at all times turn a probabilistic model right into a point-forecasting model, by taking empirical means or medians.
The Time Series Transformer
By way of modeling time series data that are sequential in nature, as one can imagine, researchers have give you models which use Recurrent Neural Networks (RNN) like LSTM or GRU, or Convolutional Networks (CNN), and more recently Transformer based methods which fit naturally to the time series forecasting setting.
On this blog post, we will leverage the vanilla Transformer (Vaswani et al., 2017) for the univariate probabilistic forecasting task (i.e. predicting every time series’ 1-d distribution individually). The Encoder-Decoder Transformer is a natural alternative for forecasting because it encapsulates several inductive biases nicely.
To start with, using an Encoder-Decoder architecture is useful at inference time, where typically for some logged data we want to forecast some prediction steps into the longer term. This could be considered analogous to the text generation task where given some context, we sample the subsequent token and pass it back into the decoder (also called “autoregressive generation”). Similarly here we can even, given some distribution type, sample from it to supply forecasts up until our desired prediction horizon. That is often known as Ancestral Sampling. Here is an awesome blog post about sampling within the context of language models.
Secondly, a Transformer helps us to coach on time series data which could contain hundreds of time points. It may not be feasible to input all the history of a time series without delay to the model, because of the time- and memory constraints of the eye mechanism. Thus, one can consider some appropriate context window and sample this window and the next prediction length sized window from the training data when constructing batches for stochastic gradient descent (SGD). The context sized window could be passed to the encoder and the prediction window to a causal-masked decoder. Because of this the decoder can only have a look at previous time steps when learning the subsequent value. That is such as how one would train a vanilla Transformer for machine translation, known as “teacher forcing”.
One other good thing about Transformers over the opposite architectures is that we will incorporate missing values (that are common within the time series setting) as an extra mask to the encoder or decoder and still train without resorting to in-filling or imputation. That is such as the attention_mask of models like BERT and GPT-2 within the Transformers library, to not include padding tokens within the computation of the eye matrix.
A drawback of the Transformer architecture is the limit to the sizes of the context and prediction windows due to the quadratic compute and memory requirements of the vanilla Transformer, see Tay et al., 2020. Moreover, because the Transformer is a strong architecture, it’d overfit or learn spurious correlations rather more easily in comparison with other methods.
The 🤗 Transformers library comes with a vanilla probabilistic time series Transformer model, simply called the Time Series Transformer. Within the sections below, we’ll show tips on how to train such a model on a custom dataset.
Set-up Environment
First, let’s install the mandatory libraries: 🤗 Transformers, 🤗 Datasets, 🤗 Evaluate, 🤗 Speed up and GluonTS.
As we are going to show, GluonTS will likely be used for transforming the info to create features in addition to for creating appropriate training, validation and test batches.
!pip install -q transformers
!pip install -q datasets
!pip install -q evaluate
!pip install -q speed up
!pip install -q gluonts ujson
Load Dataset
On this blog post, we’ll use the tourism_monthly dataset, which is out there on the Hugging Face Hub. This dataset comprises monthly tourism volumes for 366 regions in Australia.
This dataset is a component of the Monash Time Series Forecasting repository, a set of time series datasets from plenty of domains. It might probably be viewed because the GLUE benchmark of time series forecasting.
from datasets import load_dataset
dataset = load_dataset("monash_tsf", "tourism_monthly")
As could be seen, the dataset comprises 3 splits: train, validation and test.
dataset
>>> DatasetDict({
train: Dataset({
features: ['start', 'target', 'feat_static_cat', 'feat_dynamic_real', 'item_id'],
num_rows: 366
})
test: Dataset({
features: ['start', 'target', 'feat_static_cat', 'feat_dynamic_real', 'item_id'],
num_rows: 366
})
validation: Dataset({
features: ['start', 'target', 'feat_static_cat', 'feat_dynamic_real', 'item_id'],
num_rows: 366
})
})
Each example comprises just a few keys, of which start and goal are a very powerful ones. Allow us to have a have a look at the primary time series within the dataset:
train_example = dataset['train'][0]
train_example.keys()
>>> dict_keys(['start', 'target', 'feat_static_cat', 'feat_dynamic_real', 'item_id'])
The start simply indicates the beginning of the time series (as a datetime), and the goal comprises the actual values of the time series.
The start will likely be useful so as to add time related features to the time series values, as extra input to the model (equivalent to “month of yr”). Since we all know the frequency of the info is monthly, we all know as an example that the second value has the timestamp 1979-02-01, etc.
print(train_example['start'])
print(train_example['target'])
>>> 1979-01-01 00:00:00
[1149.8699951171875, 1053.8001708984375, ..., 5772.876953125]
The validation set comprises the identical data because the training set, only for a prediction_length longer period of time. This permits us to validate the model’s predictions against the bottom truth.
The test set is again one prediction_length longer data in comparison with the validation set (or some multiple of prediction_length longer data in comparison with the training set for testing on multiple rolling windows).
validation_example = dataset['validation'][0]
validation_example.keys()
>>> dict_keys(['start', 'target', 'feat_static_cat', 'feat_dynamic_real', 'item_id'])
The initial values are the exact same because the corresponding training example:
print(validation_example['start'])
print(validation_example['target'])
>>> 1979-01-01 00:00:00
[1149.8699951171875, 1053.8001708984375, ..., 5985.830078125]
Nevertheless, this instance has prediction_length=24 additional values in comparison with the training example. Allow us to confirm it.
freq = "1M"
prediction_length = 24
assert len(train_example["target"]) + prediction_length == len(
validation_example["target"]
)
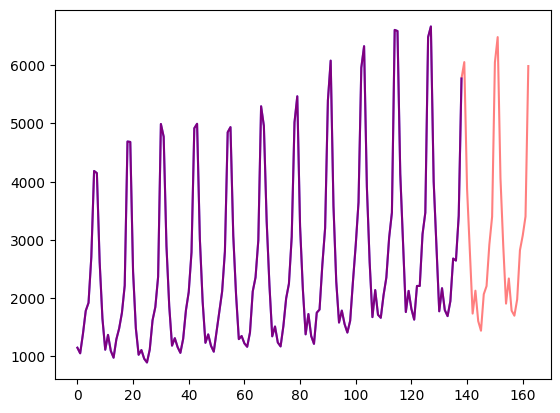
Let’s visualize this:
import matplotlib.pyplot as plt
figure, axes = plt.subplots()
axes.plot(train_example["target"], color="blue")
axes.plot(validation_example["target"], color="red", alpha=0.5)
plt.show()
Let’s split up the info:
train_dataset = dataset["train"]
test_dataset = dataset["test"]
Update start to pd.Period
The very first thing we’ll do is convert the start feature of every time series to a pandas Period index using the info’s freq:
from functools import lru_cache
import pandas as pd
import numpy as np
@lru_cache(10_000)
def convert_to_pandas_period(date, freq):
return pd.Period(date, freq)
def transform_start_field(batch, freq):
batch["start"] = [convert_to_pandas_period(date, freq) for date in batch["start"]]
return batch
We now use datasets‘ set_transform functionality to do that on-the-fly in place:
from functools import partial
train_dataset.set_transform(partial(transform_start_field, freq=freq))
test_dataset.set_transform(partial(transform_start_field, freq=freq))
Define the Model
Next, let’s instantiate a model. The model will likely be trained from scratch, hence we can’t use the from_pretrained method here, but somewhat randomly initialize the model from a config.
We specify a few additional parameters to the model:
prediction_length(in our case,24months): that is the horizon that the decoder of the Transformer will learn to predict for;context_length: the model will set thecontext_length(input of the encoder) equal to theprediction_length, if nocontext_lengthis specified;lagsfor a given frequency: these specify how much we “look back”, to be added as additional features. e.g. for aDay by dayfrequency we’d consider a glance back of[1, 2, 7, 30, ...]or in other words look back 1, 2, … days while forMinutedata we’d consider[1, 30, 60, 60*24, ...]etc.;- the variety of time features: in our case, this will likely be
2as we’ll addMonthOfYearandAgefeatures; - the variety of static categorical features: in our case, this will likely be just
1as we’ll add a single “time series ID” feature; - the cardinality: the variety of values of every static categorical feature, as a listing which for our case will likely be
[366]as now we have 366 different time series - the embedding dimension: the embedding dimension for every static categorical feature, as a listing, for instance
[3]means the model will learn an embedding vector of size3for every of the366time series (regions).
Let’s use the default lags provided by GluonTS for the given frequency (“monthly”):
from gluonts.time_feature import get_lags_for_frequency
lags_sequence = get_lags_for_frequency(freq)
print(lags_sequence)
>>> [1, 2, 3, 4, 5, 6, 7, 11, 12, 13, 23, 24, 25, 35, 36, 37]
Because of this we’ll look back as much as 37 months for every time step, as additional features.
Let’s also check the default time features that GluonTS provides us:
from gluonts.time_feature import time_features_from_frequency_str
time_features = time_features_from_frequency_str(freq)
print(time_features)
>>> [0x7fa496d0ca70>]
On this case, there’s only a single feature, namely “month of yr”. Because of this for every time step, we’ll add the month as a scalar value (e.g. 1 in case the timestamp is “january”, 2 in case the timestamp is “february”, etc.).
We now have all the pieces to define the model:
from transformers import TimeSeriesTransformerConfig, TimeSeriesTransformerForPrediction
config = TimeSeriesTransformerConfig(
prediction_length=prediction_length,
context_length=prediction_length * 2,
lags_sequence=lags_sequence,
num_time_features=len(time_features) + 1,
num_static_categorical_features=1,
cardinality=[len(train_dataset)],
embedding_dimension=[2],
encoder_layers=4,
decoder_layers=4,
d_model=32,
)
model = TimeSeriesTransformerForPrediction(config)
Note that, much like other models within the 🤗 Transformers library, TimeSeriesTransformerModel corresponds to the encoder-decoder Transformer with none head on top, and TimeSeriesTransformerForPrediction corresponds to TimeSeriesTransformerModel with a distribution head on top. By default, the model uses a Student-t distribution (but that is configurable):
model.config.distribution_output
>>> student_t
That is a vital difference with Transformers for NLP, where the pinnacle typically consists of a hard and fast categorical distribution implemented as an nn.Linear layer.
Define Transformations
Next, we define the transformations for the info, particularly for the creation of the time features (based on the dataset or universal ones).
Again, we’ll use the GluonTS library for this. We define a Chain of transformations (which is a bit comparable to torchvision.transforms.Compose for images). It allows us to mix several transformations right into a single pipeline.
from gluonts.time_feature import (
time_features_from_frequency_str,
TimeFeature,
get_lags_for_frequency,
)
from gluonts.dataset.field_names import FieldName
from gluonts.transform import (
AddAgeFeature,
AddObservedValuesIndicator,
AddTimeFeatures,
AsNumpyArray,
Chain,
ExpectedNumInstanceSampler,
InstanceSplitter,
RemoveFields,
SelectFields,
SetField,
TestSplitSampler,
Transformation,
ValidationSplitSampler,
VstackFeatures,
RenameFields,
)
The transformations below are annotated with comments, to clarify what they do. At a high level, we are going to iterate over the person time series of our dataset and add/remove fields or features:
from transformers import PretrainedConfig
def create_transformation(freq: str, config: PretrainedConfig) -> Transformation:
remove_field_names = []
if config.num_static_real_features == 0:
remove_field_names.append(FieldName.FEAT_STATIC_REAL)
if config.num_dynamic_real_features == 0:
remove_field_names.append(FieldName.FEAT_DYNAMIC_REAL)
if config.num_static_categorical_features == 0:
remove_field_names.append(FieldName.FEAT_STATIC_CAT)
return Chain(
[RemoveFields(field_names=remove_field_names)]
+ (
[
AsNumpyArray(
field=FieldName.FEAT_STATIC_CAT,
expected_ndim=1,
dtype=int,
)
]
if config.num_static_categorical_features > 0
else []
)
+ (
[
AsNumpyArray(
field=FieldName.FEAT_STATIC_REAL,
expected_ndim=1,
)
]
if config.num_static_real_features > 0
else []
)
+ [
AsNumpyArray(
field=FieldName.TARGET,
expected_ndim=1 if config.input_size == 1 else 2,
),
AddObservedValuesIndicator(
target_field=FieldName.TARGET,
output_field=FieldName.OBSERVED_VALUES,
),
AddTimeFeatures(
start_field=FieldName.START,
target_field=FieldName.TARGET,
output_field=FieldName.FEAT_TIME,
time_features=time_features_from_frequency_str(freq),
pred_length=config.prediction_length,
),
AddAgeFeature(
target_field=FieldName.TARGET,
output_field=FieldName.FEAT_AGE,
pred_length=config.prediction_length,
log_scale=True,
),
VstackFeatures(
output_field=FieldName.FEAT_TIME,
input_fields=[FieldName.FEAT_TIME, FieldName.FEAT_AGE]
+ (
[FieldName.FEAT_DYNAMIC_REAL]
if config.num_dynamic_real_features > 0
else []
),
),
RenameFields(
mapping={
FieldName.FEAT_STATIC_CAT: "static_categorical_features",
FieldName.FEAT_STATIC_REAL: "static_real_features",
FieldName.FEAT_TIME: "time_features",
FieldName.TARGET: "values",
FieldName.OBSERVED_VALUES: "observed_mask",
}
),
]
)
Define InstanceSplitter
For training/validation/testing we next create an InstanceSplitter which is used to sample windows from the dataset (as, remember, we won’t pass the whole history of values to the Transformer because of time- and memory constraints).
The instance splitter samples random context_length sized and subsequent prediction_length sized windows from the info, and appends a past_ or future_ key to any temporal keys in time_series_fields for the respective windows. The instance splitter could be configured into three different modes:
mode="train": Here we sample the context and prediction length windows randomly from the dataset given to it (the training dataset)mode="validation": Here we sample the very last context length window and prediction window from the dataset given to it (for the back-testing or validation likelihood calculations)mode="test": Here we sample the very last context length window only (for the prediction use case)
from gluonts.transform.sampler import InstanceSampler
from typing import Optional
def create_instance_splitter(
config: PretrainedConfig,
mode: str,
train_sampler: Optional[InstanceSampler] = None,
validation_sampler: Optional[InstanceSampler] = None,
) -> Transformation:
assert mode in ["train", "validation", "test"]
instance_sampler = {
"train": train_sampler
or ExpectedNumInstanceSampler(
num_instances=1.0, min_future=config.prediction_length
),
"validation": validation_sampler
or ValidationSplitSampler(min_future=config.prediction_length),
"test": TestSplitSampler(),
}[mode]
return InstanceSplitter(
target_field="values",
is_pad_field=FieldName.IS_PAD,
start_field=FieldName.START,
forecast_start_field=FieldName.FORECAST_START,
instance_sampler=instance_sampler,
past_length=config.context_length + max(config.lags_sequence),
future_length=config.prediction_length,
time_series_fields=["time_features", "observed_mask"],
)
Create DataLoaders
Next, it is time to create the DataLoaders, which permit us to have batches of (input, output) pairs – or in other words (past_values, future_values).
from typing import Iterable
import torch
from gluonts.itertools import Cached, Cyclic
from gluonts.dataset.loader import as_stacked_batches
def create_train_dataloader(
config: PretrainedConfig,
freq,
data,
batch_size: int,
num_batches_per_epoch: int,
shuffle_buffer_length: Optional[int] = None,
cache_data: bool = True,
**kwargs,
) -> Iterable:
PREDICTION_INPUT_NAMES = [
"past_time_features",
"past_values",
"past_observed_mask",
"future_time_features",
]
if config.num_static_categorical_features > 0:
PREDICTION_INPUT_NAMES.append("static_categorical_features")
if config.num_static_real_features > 0:
PREDICTION_INPUT_NAMES.append("static_real_features")
TRAINING_INPUT_NAMES = PREDICTION_INPUT_NAMES + [
"future_values",
"future_observed_mask",
]
transformation = create_transformation(freq, config)
transformed_data = transformation.apply(data, is_train=True)
if cache_data:
transformed_data = Cached(transformed_data)
instance_splitter = create_instance_splitter(config, "train")
stream = Cyclic(transformed_data).stream()
training_instances = instance_splitter.apply(stream)
return as_stacked_batches(
training_instances,
batch_size=batch_size,
shuffle_buffer_length=shuffle_buffer_length,
field_names=TRAINING_INPUT_NAMES,
output_type=torch.tensor,
num_batches_per_epoch=num_batches_per_epoch,
)
def create_backtest_dataloader(
config: PretrainedConfig,
freq,
data,
batch_size: int,
**kwargs,
):
PREDICTION_INPUT_NAMES = [
"past_time_features",
"past_values",
"past_observed_mask",
"future_time_features",
]
if config.num_static_categorical_features > 0:
PREDICTION_INPUT_NAMES.append("static_categorical_features")
if config.num_static_real_features > 0:
PREDICTION_INPUT_NAMES.append("static_real_features")
transformation = create_transformation(freq, config)
transformed_data = transformation.apply(data)
instance_sampler = create_instance_splitter(config, "validation")
testing_instances = instance_sampler.apply(transformed_data, is_train=True)
return as_stacked_batches(
testing_instances,
batch_size=batch_size,
output_type=torch.tensor,
field_names=PREDICTION_INPUT_NAMES,
)
We’ve a test dataloader helper for completion, though we is not going to use it here. This is helpful in a production setting where we wish to begin forecasting from the tip of a given time series. Thus, the test dataloader will sample the very last context window from the dataset provided and pass it to the model.
def create_test_dataloader(
config: PretrainedConfig,
freq,
data,
batch_size: int,
**kwargs,
):
PREDICTION_INPUT_NAMES = [
"past_time_features",
"past_values",
"past_observed_mask",
"future_time_features",
]
if config.num_static_categorical_features > 0:
PREDICTION_INPUT_NAMES.append("static_categorical_features")
if config.num_static_real_features > 0:
PREDICTION_INPUT_NAMES.append("static_real_features")
transformation = create_transformation(freq, config)
transformed_data = transformation.apply(data, is_train=False)
instance_sampler = create_instance_splitter(config, "test")
testing_instances = instance_sampler.apply(transformed_data, is_train=False)
return as_stacked_batches(
testing_instances,
batch_size=batch_size,
output_type=torch.tensor,
field_names=PREDICTION_INPUT_NAMES,
)
train_dataloader = create_train_dataloader(
config=config,
freq=freq,
data=train_dataset,
batch_size=256,
num_batches_per_epoch=100,
)
test_dataloader = create_backtest_dataloader(
config=config,
freq=freq,
data=test_dataset,
batch_size=64,
)
Let’s check the primary batch:
batch = next(iter(train_dataloader))
for k, v in batch.items():
print(k, v.shape, v.type())
>>> past_time_features torch.Size([256, 85, 2]) torch.FloatTensor
past_values torch.Size([256, 85]) torch.FloatTensor
past_observed_mask torch.Size([256, 85]) torch.FloatTensor
future_time_features torch.Size([256, 24, 2]) torch.FloatTensor
static_categorical_features torch.Size([256, 1]) torch.LongTensor
future_values torch.Size([256, 24]) torch.FloatTensor
future_observed_mask torch.Size([256, 24]) torch.FloatTensor
As could be seen, we do not feed input_ids and attention_mask to the encoder (as could be the case for NLP models), but somewhat past_values, together with past_observed_mask, past_time_features, and static_categorical_features.
The decoder inputs consist of future_values, future_observed_mask and future_time_features. The future_values could be seen because the equivalent of decoder_input_ids in NLP.
We confer with the docs for an in depth explanation for every of them.
Forward Pass
Let’s perform a single forward pass with the batch we just created:
outputs = model(
past_values=batch["past_values"],
past_time_features=batch["past_time_features"],
past_observed_mask=batch["past_observed_mask"],
static_categorical_features=batch["static_categorical_features"]
if config.num_static_categorical_features > 0
else None,
static_real_features=batch["static_real_features"]
if config.num_static_real_features > 0
else None,
future_values=batch["future_values"],
future_time_features=batch["future_time_features"],
future_observed_mask=batch["future_observed_mask"],
output_hidden_states=True,
)
print("Loss:", outputs.loss.item())
>>> Loss: 9.069628715515137
Note that the model is returning a loss. This is feasible because the decoder routinely shifts the future_values one position to the appropriate to be able to have the labels. This permits computing a loss between the anticipated values and the labels.
Also, note that the decoder uses a causal mask to not look into the longer term because the values it must predict are within the future_values tensor.
Train the Model
It is time to train the model! We’ll use a normal PyTorch training loop.
We are going to use the 🤗 Speed up library here, which routinely places the model, optimizer and dataloader on the suitable device.
from speed up import Accelerator
from torch.optim import AdamW
accelerator = Accelerator()
device = accelerator.device
model.to(device)
optimizer = AdamW(model.parameters(), lr=6e-4, betas=(0.9, 0.95), weight_decay=1e-1)
model, optimizer, train_dataloader = accelerator.prepare(
model,
optimizer,
train_dataloader,
)
model.train()
for epoch in range(40):
for idx, batch in enumerate(train_dataloader):
optimizer.zero_grad()
outputs = model(
static_categorical_features=batch["static_categorical_features"].to(device)
if config.num_static_categorical_features > 0
else None,
static_real_features=batch["static_real_features"].to(device)
if config.num_static_real_features > 0
else None,
past_time_features=batch["past_time_features"].to(device),
past_values=batch["past_values"].to(device),
future_time_features=batch["future_time_features"].to(device),
future_values=batch["future_values"].to(device),
past_observed_mask=batch["past_observed_mask"].to(device),
future_observed_mask=batch["future_observed_mask"].to(device),
)
loss = outputs.loss
accelerator.backward(loss)
optimizer.step()
if idx % 100 == 0:
print(loss.item())
Inference
At inference time, it’s advisable to make use of the generate() method for autoregressive generation, much like NLP models.
Forecasting involves getting data from the test instance sampler, which is able to sample the very last context_length sized window of values from every time series within the dataset, and pass it to the model. Note that we pass future_time_features, that are known ahead of time, to the decoder.
The model will autoregressively sample a certain variety of values from the anticipated distribution and pass them back to the decoder to return the prediction outputs:
model.eval()
forecasts = []
for batch in test_dataloader:
outputs = model.generate(
static_categorical_features=batch["static_categorical_features"].to(device)
if config.num_static_categorical_features > 0
else None,
static_real_features=batch["static_real_features"].to(device)
if config.num_static_real_features > 0
else None,
past_time_features=batch["past_time_features"].to(device),
past_values=batch["past_values"].to(device),
future_time_features=batch["future_time_features"].to(device),
past_observed_mask=batch["past_observed_mask"].to(device),
)
forecasts.append(outputs.sequences.cpu().numpy())
The model outputs a tensor of shape (batch_size, variety of samples, prediction length).
On this case, we get 100 possible values for the subsequent 24 months (for every example within the batch which is of size 64):
forecasts[0].shape
>>> (64, 100, 24)
We’ll stack them vertically, to get forecasts for all time-series within the test dataset:
forecasts = np.vstack(forecasts)
print(forecasts.shape)
>>> (366, 100, 24)
We will evaluate the resulting forecast with respect to the bottom truth out of sample values present within the test set. We are going to use the MASE and sMAPE metrics which we calculate for every time series within the dataset:
from evaluate import load
from gluonts.time_feature import get_seasonality
mase_metric = load("evaluate-metric/mase")
smape_metric = load("evaluate-metric/smape")
forecast_median = np.median(forecasts, 1)
mase_metrics = []
smape_metrics = []
for item_id, ts in enumerate(test_dataset):
training_data = ts["target"][:-prediction_length]
ground_truth = ts["target"][-prediction_length:]
mase = mase_metric.compute(
predictions=forecast_median[item_id],
references=np.array(ground_truth),
training=np.array(training_data),
periodicity=get_seasonality(freq))
mase_metrics.append(mase["mase"])
smape = smape_metric.compute(
predictions=forecast_median[item_id],
references=np.array(ground_truth),
)
smape_metrics.append(smape["smape"])
print(f"MASE: {np.mean(mase_metrics)}")
>>> MASE: 1.2564196892177717
print(f"sMAPE: {np.mean(smape_metrics)}")
>>> sMAPE: 0.1609541520852549
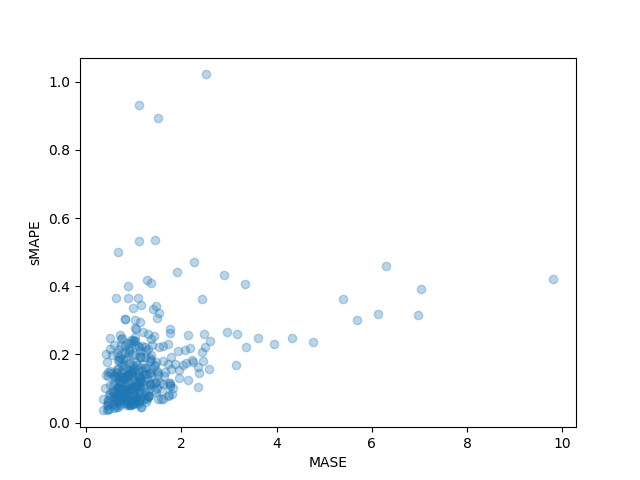
We can even plot the person metrics of every time series within the dataset and observe that a handful of time series contribute so much to the ultimate test metric:
plt.scatter(mase_metrics, smape_metrics, alpha=0.3)
plt.xlabel("MASE")
plt.ylabel("sMAPE")
plt.show()
To plot the prediction for any time series with respect the bottom truth test data we define the next helper:
import matplotlib.dates as mdates
def plot(ts_index):
fig, ax = plt.subplots()
index = pd.period_range(
start=test_dataset[ts_index][FieldName.START],
periods=len(test_dataset[ts_index][FieldName.TARGET]),
freq=freq,
).to_timestamp()
ax.xaxis.set_major_locator(mdates.MonthLocator(bymonth=(1, 7)))
ax.xaxis.set_minor_locator(mdates.MonthLocator())
ax.plot(
index[-2*prediction_length:],
test_dataset[ts_index]["target"][-2*prediction_length:],
label="actual",
)
plt.plot(
index[-prediction_length:],
np.median(forecasts[ts_index], axis=0),
label="median",
)
plt.fill_between(
index[-prediction_length:],
forecasts[ts_index].mean(0) - forecasts[ts_index].std(axis=0),
forecasts[ts_index].mean(0) + forecasts[ts_index].std(axis=0),
alpha=0.3,
interpolate=True,
label="+/- 1-std",
)
plt.legend()
plt.show()
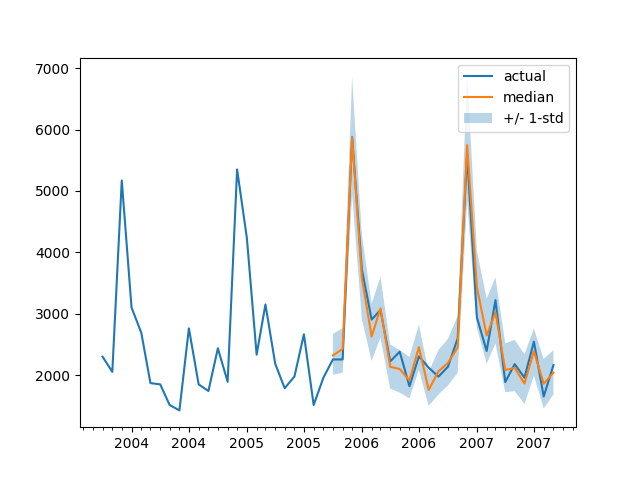
For instance:
plot(334)
How will we compare against other models? The Monash Time Series Repository has a comparison table of test set MASE metrics which we will add to:
| Dataset | SES | Theta | TBATS | ETS | (DHR-)ARIMA | PR | CatBoost | FFNN | DeepAR | N-BEATS | WaveNet | Transformer (Our) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Tourism Monthly | 3.306 | 1.649 | 1.751 | 1.526 | 1.589 | 1.678 | 1.699 | 1.582 | 1.409 | 1.574 | 1.482 | 1.256 |
Note that, with our model, we’re beating all other models reported (see also table 2 within the corresponding paper), and we didn’t do any hyperparameter tuning. We just trained the Transformer for 40 epochs.
In fact, we should be careful with just claiming state-of-the-art results on time series with neural networks, because it seems “XGBoost is usually all you would like”. We are only very curious to see how far neural networks can bring us, and whether Transformers are going to be useful on this domain. This particular dataset seems to point that it’s definitely value exploring.
Next Steps
We’d encourage the readers to check out the notebook with other time series datasets from the Hub and replace the suitable frequency and prediction length parameters. In your datasets, one would wish to convert them to the convention utilized by GluonTS, which is explained nicely of their documentation here. We’ve also prepared an example notebook showing you tips on how to convert your dataset into the 🤗 datasets format here.
As time series researchers will know, there was numerous interest in applying Transformer based models to the time series problem. The vanilla Transformer is just one in all many attention-based models and so there’s a have to add more models to the library.
In the mean time nothing is stopping us from modeling multivariate time series, nonetheless for that one would wish to instantiate the model with a multivariate distribution head. Currently, diagonal independent distributions are supported, and other multivariate distributions will likely be added. Stay tuned for a future blog post that may include a tutorial.
One other thing on the roadmap is time series classification. This entails adding a time series model with a classification head to the library, for the anomaly detection task for instance.
The present model assumes the presence of a date-time along with the time series values, which could not be the case for each time series within the wild. See as an example neuroscience datasets just like the one from WOODS. Thus, one would wish to generalize the present model to make some inputs optional in the entire pipeline.
Finally, the NLP/Vision domain has benefitted tremendously from large pre-trained models, while this shouldn’t be the case so far as we’re aware for the time series domain. Transformer based models appear to be the plain alternative in pursuing this avenue of research and we cannot wait to see what researchers and practitioners give you!